ASAFind
ASAFind is a software that predicts the intracellular location of proteins in cells with four membrane-bound complex plastids of red algal origin [1]. These plastids evolved via eukaryote-eukaryote endosymbiosis and for example can be found in diatoms and cryptophytes. The plastids in these groups of algae are not only surrounded by additional membranes compared to primary plastids (as found in red-, green- and glaucophyte algae), but also by an additional compartment between the second and third membrane, which is referred to as the periplastidic compartment (ppc, Figure 1). The ppc is thought to correspond to the former cytosol of the eukaryotic endosymbiont and plays a role in the cellular metabolism [2,3]. Nucleus encoded plastid proteins are transported to the plastids via the endoplasmic reticulum (ER), and in this process also pass through the ppc. The N-terminal part of the pre-protein of nucleus-encoded diatom plastid proteins is an ER-type signal peptide, which is followed by a transit peptide, and the cleavage site between these two targeting domains is characterized by a characteristic sequence motif (“ASA-FAP”), such targeting pre-sequences are commonly referred to as bipartite targeting signals (Figure 1).
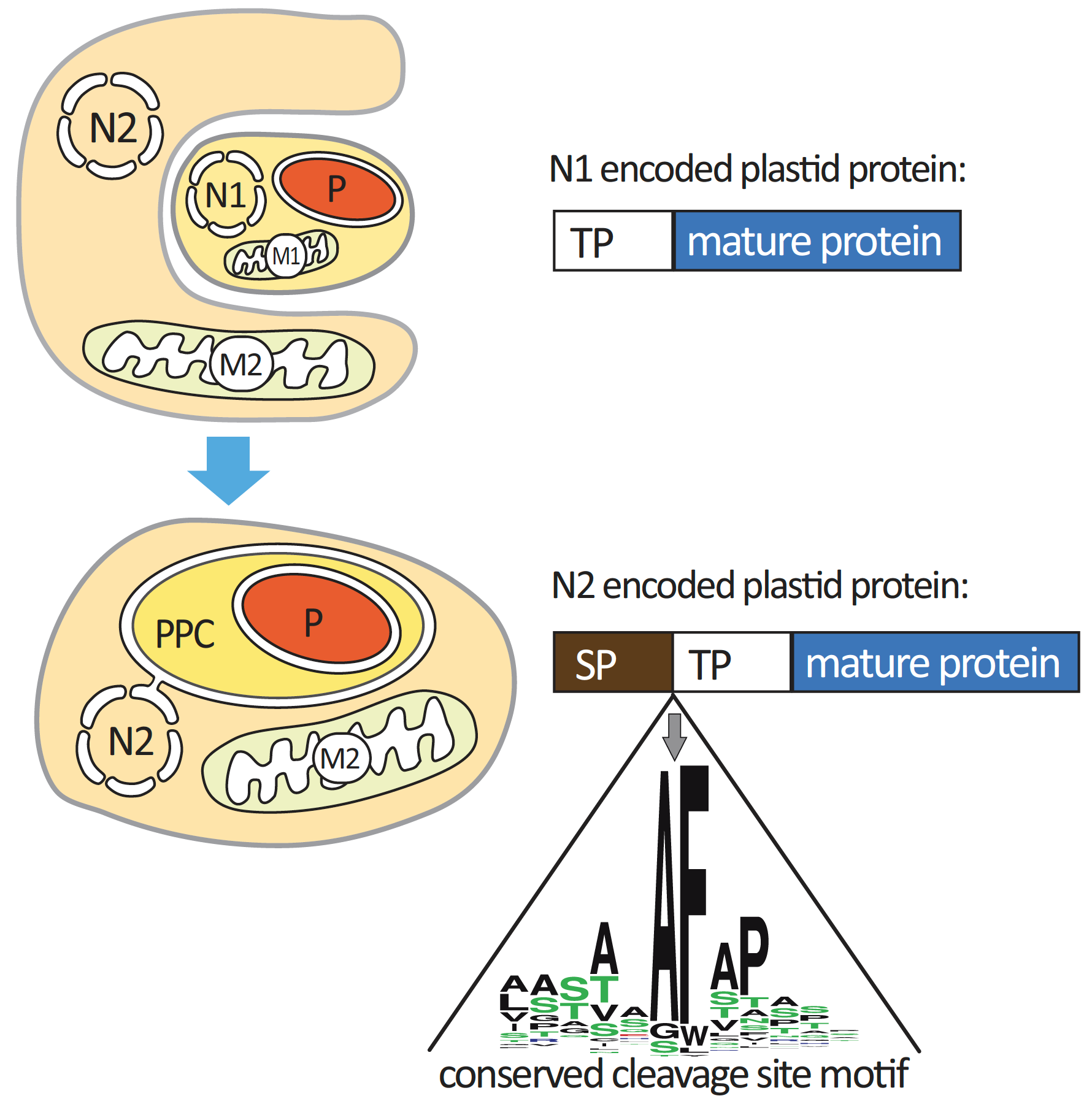
Figure 1: Complex plastid evolution and protein targeting; plastid evolution via eukaryote-eukaryote endosymbiosis leads to additional membranes and compartments surrounding the resulting plastids. In complex plastids of red algal origin, a conserved cleavage site is found at the cleavage site between signal and transit peptide domains of a bipartite targeting signal. Abbreviations: M1: host cell mitochondria, M2: symbiont mitochondria, N1: host cell nucleus, N2: symbiont nucleus, P: plastid, PPC: periplastidic compartment, SP: signal peptide, TP: transit peptide, figure reproduced from [3].
ASAFind was first developed in 2015 [1], and updated in 2025 to include predictions of ppc targeted proteins, a graphical output (Figure 2), and more options for the customization of scoring matrices. This page provides a web service to run ASAFind remotely, and instructions for download and local installation. Please register on this page to access our web-service and for contacting us through the contact form (registration is not necessary for the download).
Further information can be found in the original publication of ASAFind [1], in this book chapter with an overview on the topic of protein transport to the plastid in cells with complex plastids of red algal origin [2], or in this book chapter with an overview on the evolutionary pathways and physiological consequences of complex endosymbioses [3].
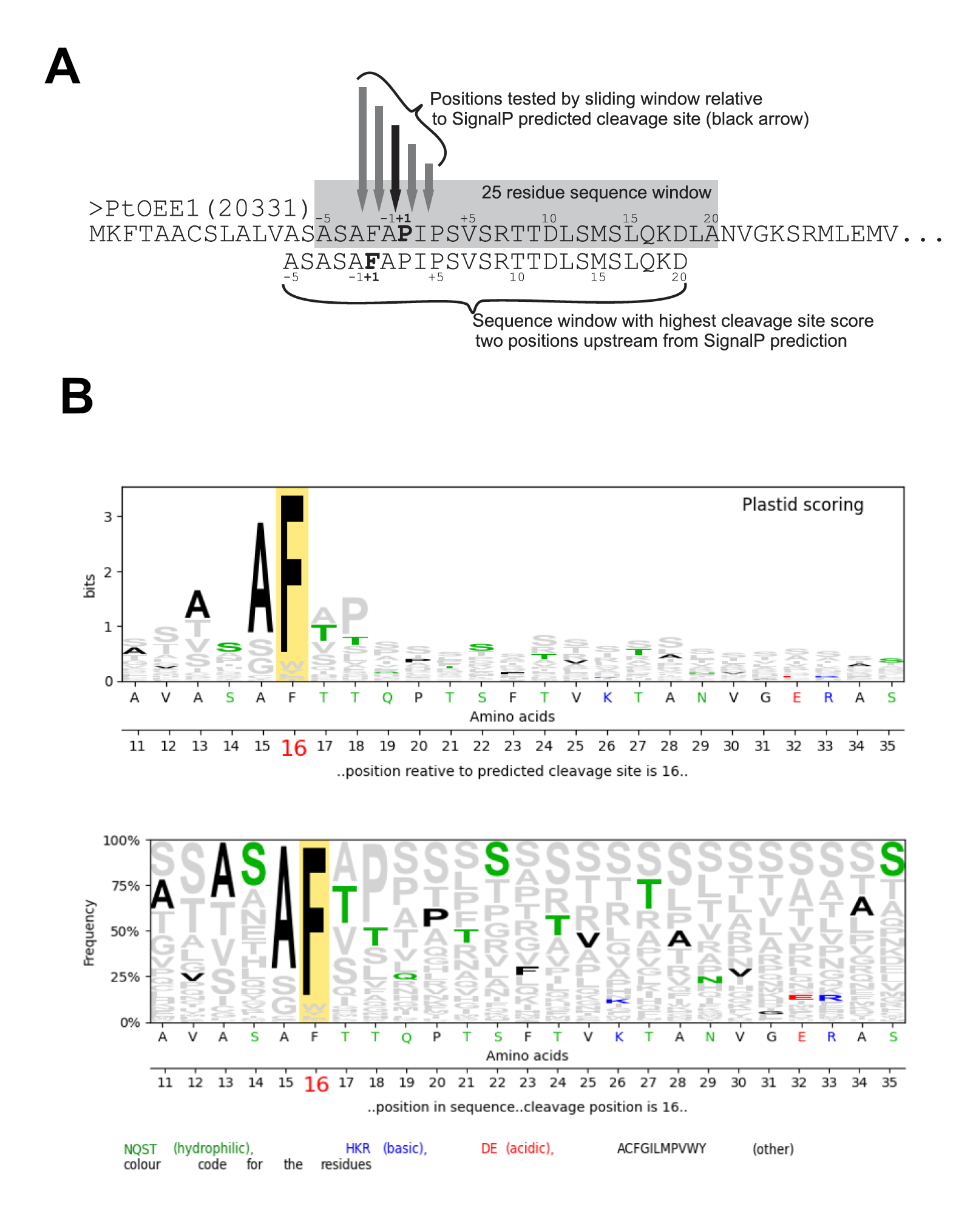
Figure 2: A ASAFind scores the input sequences according to a matric calculated from a set of known diatom plastid proteins, reproduced from [1]. B Graphical output of ASAFind 2.0; the residues of the tested sequence are highlighted against the background of the scoring matrix.
References
- Gruber, A., Rocap, G., Kroth, P. G., Armbrust, E. v & Mock, T. Plastid proteome prediction for diatoms and other algae with secondary plastids of the red lineage. Plant Journal 81, 519–528 (2015). https://dx.doi.org/10.1111/tpj.12734
- Gruber, A. & Kroth, P. G. Translocation of Proteins into Four Membrane-Bound Complex Plastids of Red Algal Origin. in Endosymbiotic Organelle Acquisition: Solutions to the Problem of Protein Localization and Membrane Passage (eds. Schwartzbach, S., Kroth, P. G. & Oborník, M.) 433–463 (Springer International Publishing, 2024). https://dx.doi.org/10.1007/978-3-031-57446-7_15
- Gruber, A. & Oborník, M. Evolution of plastids and mitochondria in diatoms. in Diatom Photosynthesis: From Primary Production to High Value (eds. Goessling, J. W., Serôdio, J. & Lavaud, J.) 81–111 (Scrivener Publishing LLC, 2024). https://dx.doi.org/10.1002/9781119842156.ch3
Acknowledgement
ASAFind 2.0 was developed with the support of ELIXIR CZ Research Infrastructure (ID LM2023055, MEYS CR); this web-service is offered by the Faculty of Science at the University of South Bohemia, with support from ELIXIR CZ.